|
Transnetyx
mouse strains Mouse Strains, supplied by Transnetyx, used in various techniques. Bioz Stars score: 97/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/mouse strains/product/Transnetyx Average 97 stars, based on 1 article reviews
mouse strains - by Bioz Stars,
2026-04
97/100 stars
|
Buy from Supplier |
|
CLS Cell Lines Service GmbH
pig kidney epithelial cell line llc pk1 Pig Kidney Epithelial Cell Line Llc Pk1, supplied by CLS Cell Lines Service GmbH, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/pig kidney epithelial cell line llc pk1/product/CLS Cell Lines Service GmbH Average 93 stars, based on 1 article reviews
pig kidney epithelial cell line llc pk1 - by Bioz Stars,
2026-04
93/100 stars
|
Buy from Supplier |
|
Molecular Devices LLC
cloneselect a g  Cloneselect A G, supplied by Molecular Devices LLC, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/cloneselect a g/product/Molecular Devices LLC Average 96 stars, based on 1 article reviews
cloneselect a g - by Bioz Stars,
2026-04
96/100 stars
|
Buy from Supplier |
|
Danaher Inc
multi well plate reader image xpress pico automated bioimager system  Multi Well Plate Reader Image Xpress Pico Automated Bioimager System, supplied by Danaher Inc, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/multi well plate reader image xpress pico automated bioimager system/product/Danaher Inc Average 96 stars, based on 1 article reviews
multi well plate reader image xpress pico automated bioimager system - by Bioz Stars,
2026-04
96/100 stars
|
Buy from Supplier |
|
Tymora Analytical Operations LLC
proteomics analysis  Proteomics Analysis, supplied by Tymora Analytical Operations LLC, used in various techniques. Bioz Stars score: 95/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/proteomics analysis/product/Tymora Analytical Operations LLC Average 95 stars, based on 1 article reviews
proteomics analysis - by Bioz Stars,
2026-04
95/100 stars
|
Buy from Supplier |
|
AllCells LLC
normal cd34+ cells  Normal Cd34+ Cells, supplied by AllCells LLC, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/normal cd34+ cells/product/AllCells LLC Average 90 stars, based on 1 article reviews
normal cd34+ cells - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Meso Scale Diagnostics LLC
akt signaling whole cell lysate kit  Akt Signaling Whole Cell Lysate Kit, supplied by Meso Scale Diagnostics LLC, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/akt signaling whole cell lysate kit/product/Meso Scale Diagnostics LLC Average 90 stars, based on 1 article reviews
akt signaling whole cell lysate kit - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
AllCells LLC
primary human cd341 hematopoietic stem cells (hscs)  Primary Human Cd341 Hematopoietic Stem Cells (Hscs), supplied by AllCells LLC, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/primary human cd341 hematopoietic stem cells (hscs)/product/AllCells LLC Average 90 stars, based on 1 article reviews
primary human cd341 hematopoietic stem cells (hscs) - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Stonehouse Enterprises LLC
cell lines  Cell Lines, supplied by Stonehouse Enterprises LLC, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/cell lines/product/Stonehouse Enterprises LLC Average 90 stars, based on 1 article reviews
cell lines - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
AllCells LLC
cd8 t cells  Cd8 T Cells, supplied by AllCells LLC, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/cd8 t cells/product/AllCells LLC Average 90 stars, based on 1 article reviews
cd8 t cells - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
Perma Pure LLC
porcine bone marrow mesenchymal stromal cells  Porcine Bone Marrow Mesenchymal Stromal Cells, supplied by Perma Pure LLC, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/porcine bone marrow mesenchymal stromal cells/product/Perma Pure LLC Average 90 stars, based on 1 article reviews
porcine bone marrow mesenchymal stromal cells - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
|
AllCells LLC
frozen healthy donor naive cd4+ t cells  Frozen Healthy Donor Naive Cd4+ T Cells, supplied by AllCells LLC, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/frozen healthy donor naive cd4+ t cells/product/AllCells LLC Average 90 stars, based on 1 article reviews
frozen healthy donor naive cd4+ t cells - by Bioz Stars,
2026-04
90/100 stars
|
Buy from Supplier |
Image Search Results

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) Conceptual diagram of retrospective clone isolation. ( B ) Different barcode-specific gRNA-dependent reporter activation circuits. CloneSelect C→T, low-copy CRISPRa, and high-copy CRISPRa. ( C ) Flow cytometry analysis of single-cell EGFP activation levels. ( D ) Barcode-dependent reporter activation of six barcoded cell lines by CloneSelect C→T. Scale bar, 50 μm. ( E ) Comparison of CloneSelect C→T, low-copy CRISPRa, and high-copy CRISPRa across the same barcode-gRNA pairs (n=3). For each approach, Welch’s t-test was performed to compare on-target (OT) and non-target (NT) activations. ( F ) CloneSelect A→G. ( G ) Comparison of CloneSelect A→G, low-copy CRISPRa, and high-copy CRISPRa across the same barcode-gRNA pairs (n=3). Welch’s t-test was performed to compare OT and NT activations. ( H ) ROC curves along varying EGFP intensity thresholds for target barcoded cells. Left, CloneSelect C→T and low-copy CRISPRa by the same targeting gRNAs. Right, CloneSelect A→G and low-copy CRISPRa for the same set of targeting gRNAs. ( I ) Performance comparison of CloneSelect C→T and CloneSelect A→G. Activated cell frequencies of OT and NT barcodes were normalized by activated cell frequencies of OT barcodes conferred by low-copy CRISPRa using the same targeting gRNA. The Mann-Whitney U test was performed to compare the two groups of datasets. * P < 0.05; ** P < 0.01; *** P < 0.001.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Isolation, Activation Assay, Flow Cytometry, MANN-WHITNEY

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A – C ) Barcode-specific gRNA-dependent activation of EGFP reporters for two barcoded HEK293T strains established for each of CloneSelect C→T ( A ), low-copy CRISPRa ( B ), and high-copy CRISPRa ( C ) (n=3). Scale bar, 50 μm. Welch’s t-test was performed to compare on-target (OT) and non-target (NT) activations. ( D ) Median EGFP intensities of genome editing-activated EGFP positive cells (n=3). The Mann-Whitney U test was performed to compare two groups. ( E ) Comparison of Target-AID variants and a nCas9 (D10A) control in the CloneSelect C→T reporter activation for the same set of barcode-gRNA pairs (n=1). Welch’s t-test was performed to compare OT and NT activations. ( F ) Reporter activation in HeLa cells by CloneSelect C→T (n=3). Welch’s t-test was performed to compare OT and NT activations. Scale bar, 80 μm. * P < 0.05; ** P < 0.01; *** P < 0.001.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Activation Assay, MANN-WHITNEY

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A and B ) Barcode-specific gRNA-dependent reporter activation of six barcoded cell lines prepared for each of low-copy CRISPRa and high-copy CRISPRa. Scale bar, 50 μm. ( C ) Median EGFP intensities of genome editing-activated EGFP positive cells (n=3). ( D ) ROC curves along varying reporter intensity thresholds for target barcoded cells. CloneSelect C→T, low-copy CRISPRa, and high-copy CRISPRa were examined for the common set of six barcodes. The Mann-Whitney U test was performed to compare two groups (* P < 0.05; ** P < 0.01; *** P < 0.001). ( E ) Frequencies of CloneSelect C→T reporter-activated cells obtained by transfection of different DNA amounts of barcode-targeting genome editing reagents. ( F ) ROC curve for each input DNA amount along varying reporter intensity thresholds for target barcoded cells.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Activation Assay, MANN-WHITNEY, Transfection

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) Different mCherry reporter variants tested to establish CloneSelect C→T. ( B and C ) mCherry expression from the different reporter variants with the first codon as GTG or ATG. Scale bar, 50 μm. ( D ) Activation of the M1V (GTG)+Δ2-9 mutant reporter with OT and NT gRNAs (n=3). Scale bar, 100 μm.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Expressing, Activation Assay, Mutagenesis

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A – C ) Barcodespecific gRNA-dependent reporter activation of three barcoded cell lines prepared for each of CloneSelect A→G, low-copy CRISPRa, and high-copy CRISPRa. Scale bar, 50 μm. ( D – F ) Flow cytometry analysis of single-cell EGFP activation levels. ( G ) Median EGFP intensities of genome editing-activated EGFP positive cells (n=3). The Mann-Whitney U test was performed to compare two groups (* P < 0.05; ** P < 0.01; *** P < 0.001). ( H ) ROC curves along varying reporter intensity thresholds for target barcoded cells (n=3).
Article Snippet: Similar to CloneSelect C→T,
Techniques: Activation Assay, Flow Cytometry, MANN-WHITNEY

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) Nucleotide compositions of barcodes in the mammalian CloneSelect C→T plasmid mini-pool. Five barcodes that had unexpected lengths were excluded from this visualization. The full barcode sequence list can be found in Table S1. ( B ) Barcode abundances in the cell population labeled by the mini-lentiviral barcode pool of CloneSelect C→T. ( C ) gRNA-dependent labeling of target barcoded cells in a population. ( D ) Flow cytometry cell sorting of reporter-activated cells. ( E ) Barcode enrichment analysis after cell sorting of the reporter-activated cells. Each row represents the barcode enrichment profile for each target isolation assay.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Plasmid Preparation, Sequencing, Labeling, Flow Cytometry, FACS, Isolation
Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
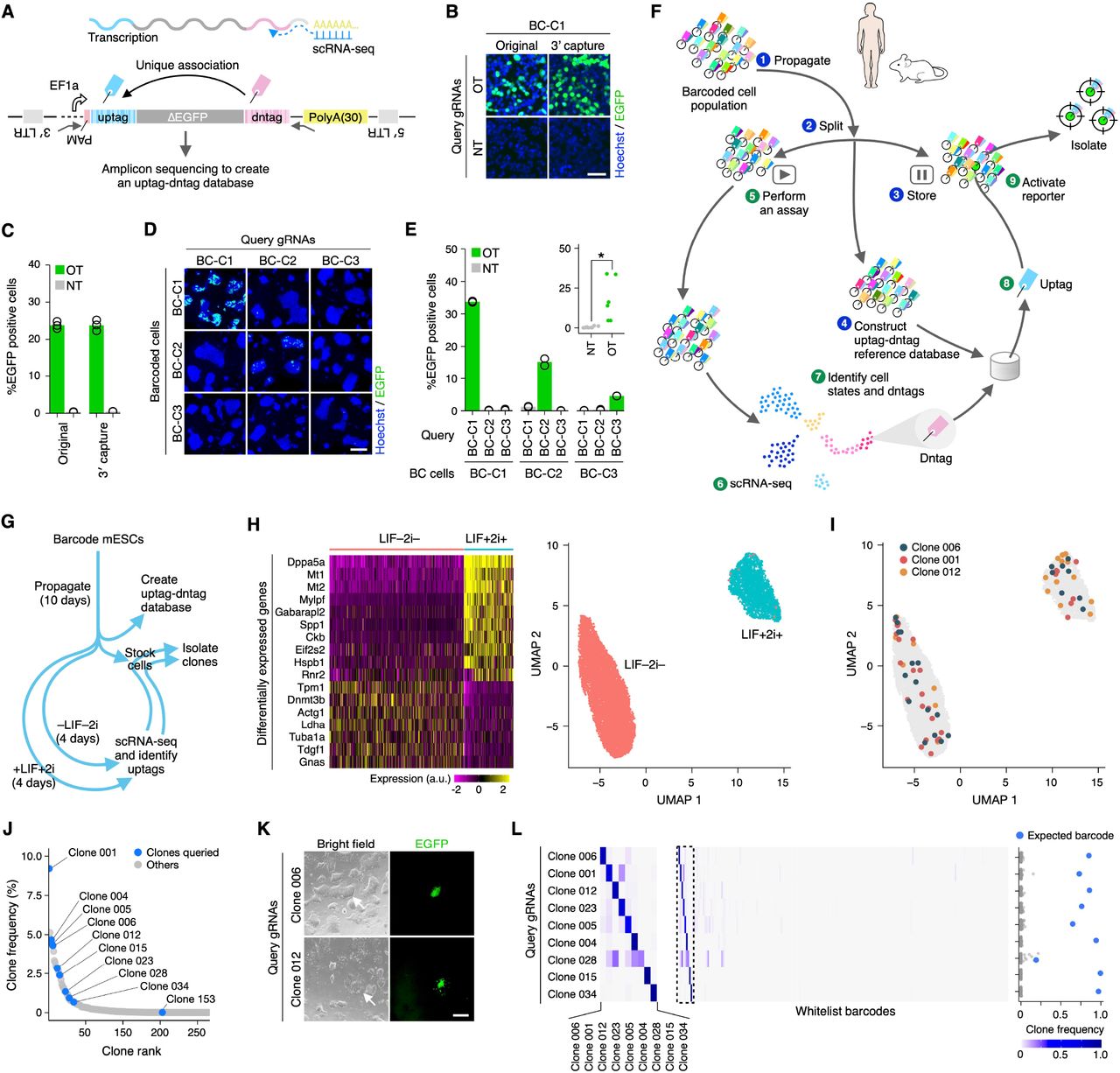
Figure Lengend Snippet: ( A ) scCloneSelect. ( B and C ) Barcode-specific gRNA-dependent reporter activation of the original CloneSelect C→T and scCloneSelect in HEK293T cells (n=3). Scale bar, 50 μm. ( D and E ) Barcode-specific gRNA-dependent reporter activation of three barcoded mESC lines by scCloneSelect. Target-AID was stably integrated prior to the barcoding. gRNAs were delivered by lentiviral transduction. Scale bar, 100 μm. Welch’s t-test was performed to compare on-target (OT) and non-target (NT) activations. * P < 0.05; ** P < 0.01; *** P < 0.001. ( F ) Schematic diagram of a scCloneSelect workflow to retrospectively isolate a cell clone demonstrating a gene expression profile of interest from a cell population stored before they demonstrate the target gene expression pattern. ( G ) mESC cell culture assays and clone isolation performed in this work. ( H ) scRNA-seq of mESC populations treated with LIF and 2i and those without LIF or 2i. ( I ) Distribution of cells for arbitrarily selected clones in the two-dimensional embedding of high-dimensional gene expression space by UMAP (uniform manifold approximation and projection). ( J ) Abundance of barcoded cell clones in the mESC population. The data was generated based on dntags identified by reamplifying the dntag reads from the original scRNA-seq libraries. ( K ) gRNA-specific activation of target barcoded clones in the mESC population. Scale bar, 50 μm. ( L ) Barcode enrichment analysis after cell sorting of the reporter-activated cells. Each row represents the barcode enrichment profile for each target isolation assay. The left heatmap was expanded from the dashed box area of the right heatmap.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Activation Assay, Stable Transfection, Transduction, Expressing, Cell Culture, Isolation, Clone Assay, Generated, FACS

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) EGFP-positive control expressions for the original CloneSelect C→T and scCloneSelect in HEK293T cells with the same genome editing conditions tested for the respective reporters (n=3). Scale bar, 50 μm. ( B ) Median EGFP intensities of base editing-activated EGFP positive cells (n=3). ( C and D ) Barcode-specific gRNA-dependent reporter activation of six barcoded cell lines by scCloneSelect (n=1). Welch’s t-test was performed to compare on-target (OT) and non-target (NT) activations (* P < 0.05; ** P < 0.01; *** P < 0.001). Scale bar, 50 μm. ( E ) RT-PCR of the scCloneSelect dntags in HEK293T. ( F ) Fraction of mESC single-cell transcriptome profiles (Drop-seq) that contained dntags and fraction of dntags reported in the uptag-dntag combination reference database. ( G ) Schematic representation of a scCloneSelect reporter activation assay where Target-AID was stably introduced to the cell population prior to barcoding and gRNA-dependent reporter activation. ( H and I ) gRNA-dependent reporter activation of target barcoded mESCs and CA1 hPSCs by scCloneSelect (n=2). Target-AID was stably integrated prior to the barcoding. Targeting gRNAs were delivered by transfection. Welch’s t-test was performed to compare OT and NT activations (* P < 0.05; ** P < 0.01; *** P < 0.001). Scale bar, 100 μm. ( J ) Schematic representation of a scCloneSelect reporter activation assay where the target gRNA and Target-AID were electroporated together to the barcoded cell population. ( K ) gRNA-dependent reporter activation of barcoded H1 hPSCs by scCloneSelect (n=2). Targeting gRNA and Target-AID were electroporated together. Scale bar, 100 μm.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Positive Control, Activation Assay, Reverse Transcription Polymerase Chain Reaction, Stable Transfection, Transfection

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) Yeast CloneSelect C→T circuit. ( B and C ) Barcode-specific gRNA-dependent reporter activation. Scale bar, 25 μm. Mean mCherry intensity measured by a plate reader was normalized by OD 595 nm (n=3). Welch’s t-test was performed to compare on-target (OT) and non-target (NT) activities. ( D ) GTG→ATG editing frequencies observed by high-throughput sequencing. Welch’s t-test was performed to compare OT and NT datasets. ( E ) Yeast colonies formed on a 10-cm agar plate after performing a target clone labeling in the yeast cell population of Pool-100. ( F–J ) Analysis of colonies isolated after clone labeling using each targeting gRNA. ( F ) mCherry positive isolates from Pool-100. ( G ) mCherry negative isolates from Pool-100. ( H ) mCherry positive isolates from Pool-1580. ( I ) mCherry negative isolates from Pool-1580. ( J ) Summary of the analysis results. * P < 0.05; ** P < 0.01; *** P < 0.001.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Activation Assay, Next-Generation Sequencing, Labeling, Isolation

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) Different mCherry reporter variants tested to establish CloneSelect C→T. The different reporter variants were tested with the first codon as GTG or ATG. Scale bar, 100 μm. ( B ) Canavanine resistance assays for different CRISPR genome editing enzymes with a gRNA targeting CAN1 gene and a control NT gRNA. For each experiment, cell concentration was normalized to 1.0 OD 595 nm and serially diluted with 10-fold increments for spotting. ( C ) Estimated CFU counts for the same assay in ( B ). ( D ) Genome editing outcomes observed by amplicon sequencing. Frequencies of mutation patterns observed across the target sequence region are shown for the same assay in ( B ). ( E ) Genome editing frequencies at the target CAN1 locus estimated by amplicon sequencing for the different enzymes. ( F ) Activation of the mCherry M1V (GTG)+Δ2-9 mutant reporter by OT and NT gRNAs. Scale bar, 200 μm. ( G ) mCherry-positive control expressions for yeast CloneSelect. Yeast cells having the positive control reporters with three different barcodes (BC-C1, BC-C2, and BC-C3) were each treated by Target-AID and three different targeting gRNAs. Scale bar, 25 μm.
Article Snippet: Similar to CloneSelect C→T,
Techniques: CRISPR, Concentration Assay, Amplification, Sequencing, Mutagenesis, Activation Assay, Positive Control

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) Bacterial CloneSelect A→G circuit. ABE and gRNA expressions were controlled by IPTG-inducible promoters, and the EGFP reporter expression was controlled by an arabinose-inducible promoter. ( B and C ) EGFP reporter activation of E. coli cells under different inducer conditions. Scale bar, 25 μm. Mean EGFP intensity measured by a plate reader was normalized by OD 595 nm (n=3). Welch’s t-test was performed to compare on-target (OT) and non-target (NT) activities. ( D ) Base editing outcomes analyzed by Sanger sequencing. ( E ) Activities of the positive control EGFP reporter under the same conditions tested for (C) (n=3). Welch’s t-test was performed to compare OT and NT activities. ( F ) Zeocin resistance marker-based circuit. ( G ) Barcode-specific gRNA-dependent Zeocin resistance reporter activation. ( H ) Schematic diagram of a bacterial CloneSelect workflow using a drug selective condition for the target barcoded cell isolation. ( I ) Abundance of barcoded cells in Pool-100 and Pool-1550. ( J ) Colonies formed on Zeocin-selective and non-selective solid agar plates after performing the reporter activation of Clone 106 in the E. coli cell population of Pool-100. ( K ) Analysis of colonies isolated from Zeocin selective and non-selective plates obtained after clone labeling using each targeting gRNA. * P < 0.05; ** P < 0.01; *** P < 0.001.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Expressing, Activation Assay, Sequencing, Positive Control, Marker, Cell Isolation, Isolation, Labeling

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) Activities of the positive control EGFP reporter. ABE and gRNA expression were controlled by an IPTG-inducible promoter, and the EGFP reporter expression was controlled by an arabinose-inducible promoter. ( B ) Base editing outcomes of the positive control reporters analyzed by Sanger sequencing. ( C ) Testing of Zeocin resistances conferred by two promoters expressing a Zeocin resistance gene with and without the upstream stop codon to block the selective marker translation. Each cell sample concentration was first adjusted to 0.1 OD 595 nm and serially diluted with 10-fold increments for spotting 5 μL. ( D ) Testing of cell viability under a non-selective condition for a constitutively active T7 promoter and the IPTG-inducible promoter to express the gRNA. OT and NT gRNAs were tested for the gRNA-dependent EGFP reporter and the positive control EGFP reporter. ABE was expressed under the IPTG-inducible promoter without IPTG provided. ( E ) gRNA-dependent Zeocin resistance reporter activation tested for the IPTG-inducible promoters with and without IPTG. ( F ) Bacterial CloneSelect using the Blasticidin resistance gene. Each cell sample concentration was first adjusted to 0.1 OD 595 nm and serially diluted with 10-fold increments for spotting 5 μL. ( G ) gRNA-dependent Blasticidin-resistance reporter activation tested for different inducer conditions and different Blasticidin concentrations. Each cell sample concentration was adjusted to 0.1 OD 595 nm for spotting 5 μL.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Positive Control, Expressing, Sequencing, Blocking Assay, Marker, Concentration Assay, Activation Assay

Journal: bioRxiv
Article Title: A multi-kingdom genetic barcoding system for precise target clone isolation
doi: 10.1101/2023.01.18.524633
Figure Lengend Snippet: ( A ) Three-gRNA-input OR gate with CloneSelect C→T that is designed to confer the EGFP reporter expression by any of the three barcode-specific gRNA-dependent GTG→ATG mutations. ( B ) Three-gRNA-input AND gate with CloneSelect A→G that is designed to confer the EGFP reporter expression when all three barcode-specific gRNA-dependent TAA→CAA mutations are provided.
Article Snippet: Similar to CloneSelect C→T,
Techniques: Expressing